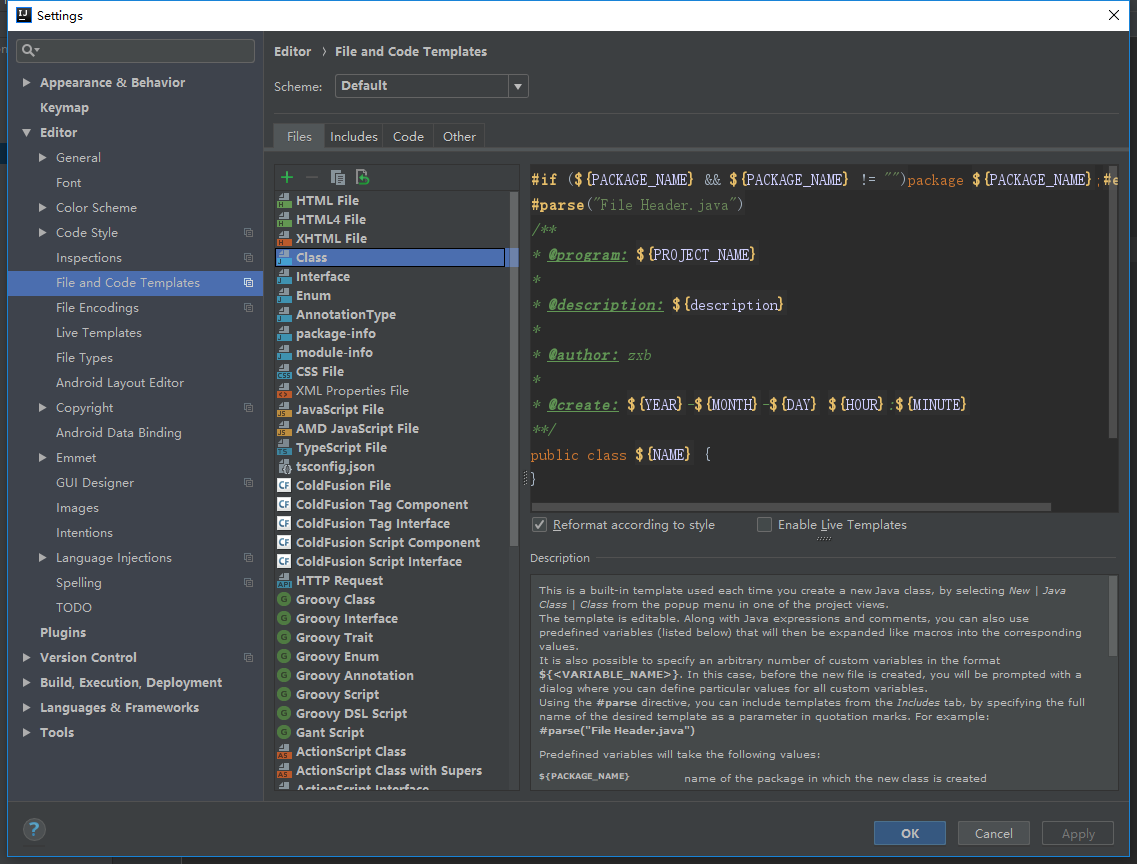
I have a x,y distribution of points for which I obtain the KDE through scipy.stats.gaussian_kde. This is my code and how the output looks (the x,y data can be obtained from here):
import numpy as np
from scipy import stats
# Obtain data from file.
data = np.loadtxt('data.dat', unpack=True)
m1, m2 = data[0], data[1]
xmin, xmax = min(m1), max(m1)
ymin, ymax = min(m2), max(m2)
# Perform a kernel density estimate (KDE) on the data
x, y = np.mgrid[xmin:xmax:100j, ymin:ymax:100j]
positions = np.vstack([x.ravel(), y.ravel()])
values = np.vstack([m1, m2])
kernel = stats.gaussian_kde(values)
f = np.reshape(kernel(positions).T, x.shape)
# Define the number that will determine the integration limits
x1, y1 = 2.5, 1.5
# Perform integration?
# Plot the results:
import matplotlib.pyplot as plt
# Set limits
plt.xlim(xmin,xmax)
plt.ylim(ymin,ymax)
# KDE density plot
plt.imshow(np.rot90(f), cmap=plt.cm.gist_earth_r, extent=[xmin, xmax, ymin, ymax])
# Draw contour lines
cset = plt.contour(x,y,f)
plt.clabel(cset, inline=1, fontsize=10)
plt.colorbar()
# Plot point
plt.scatter(x1, y1, c='r', s=35)
plt.show()

The red point with coordinates (x1, y1) has (like every point in the 2D plot) an associated value given by f (the kernel or KDE) between 0 and 0.42. Let's say that f(x1, y1) = 0.08.
I need to integrate f with integration limits in x and y given by those regions where f evaluates to less than f(x1, y1), ie: f(x, y)<0.08.
For what I've seen python can perform integration of functions and one dimensional arrays through numerical integration, but I haven't seen anything that would let me perform a numerical integration on a 2D array (the f kernel) Furthermore, I'm not sure how I would even recognize the regions given by that particular condition (ie: f(x, y)less than a given value)
Can this be done at all?
Here is a way to do it using monte carlo integration. It is a little slow, and there is randomness in the solution. The error is inversely proportional to the square root of the sample size, while the running time is directly proportional to the sample size (where sample size refers to the monte carlo sample (10000 in my example below), not the size of your data set). Here is some simple code using your kernel object.
#Compute the point below which to integrate
iso = kernel((x1,y1))
#Sample from your KDE distribution
sample = kernel.resample(size=10000)
#Filter the sample
insample = kernel(sample) < iso
#The integral you want is equivalent to the probability of drawing a point
#that gets through the filter
integral = insample.sum() / float(insample.shape[0])
print integral
I get approximately 0.2 as the answer for your data set.
A direct way is to integrate
import matplotlib.pyplot as plt
import sklearn
from scipy import integrate
import numpy as np
mean = [0, 0]
cov = [[5, 0], [0, 10]]
x, y = np.random.multivariate_normal(mean, cov, 5000).T
plt.plot(x, y, 'o')
plt.show()
sample = np.array(zip(x, y))
kde = sklearn.neighbors.KernelDensity().fit(sample)
def f_kde(x,y):
return np.exp((kde.score_samples([[x,y]])))
point = x1, y1
integrate.nquad(f_kde, [[-np.inf, x1],[-np.inf, y1]])
The problem is that, this is very slow if you do it in a large scale. For example, if you want to plot the x,y line at x (0,100), it would take a long time to calculate.
Notice: I used kde from sklearn, but I believe you can also change it into other form as well.
Using the kernel as defined in the original question:
import numpy as np
from scipy import stats
from scipy import integrate
def integ_func(kde, x1, y1):
def f_kde(x, y):
return kde((x, y))
integ = integrate.nquad(f_kde, [[-np.inf, x1], [-np.inf, y1]])
return integ
# Obtain data from file.
data = np.loadtxt('data.dat', unpack=True)
# Perform a kernel density estimate (KDE) on the data
kernel = stats.gaussian_kde(data)
# Define the number that will determine the integration limits
x1, y1 = 2.5, 1.5
print integ_func(kernel, x1, y1)