I have 10 greyscale brain MRI scans from BrainWeb. They are stored as a 4d numpy array, brains, with shape (10, 181, 217, 181). Each of the 10 brains is made up of 181 slices along the z-plane (going through the top of the head to the neck) where each slice is 181 pixels by 217 pixels in the x (ear to ear) and y (eyes to back of head) planes respectively.
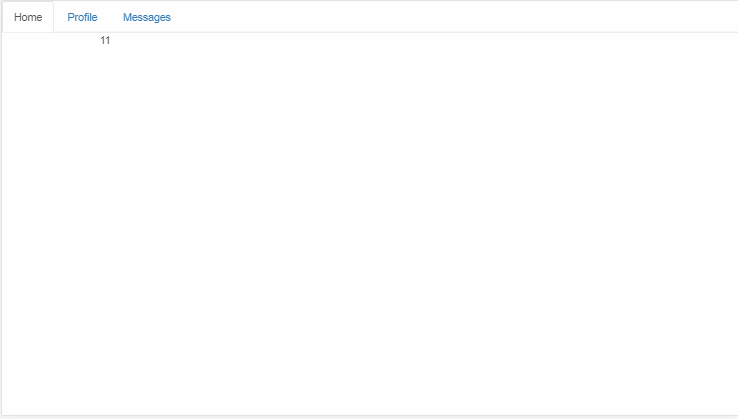
All of the brains are type dtype('float64'). The maximum pixel intensity across all brains is ~1328 and the minimum is ~0. For example, for the first brain, I calculate this by brains[0].max() giving 1328.338086605072 and brains[0].min() giving 0.0003886114541273855. Below is a plot of a slice of a brain[0]:

I want to binarize all these brain images by rescaling the pixel intensities from [0, 1328] to {0, 1}. Is my method correct?
I do this by first normalising the pixel intensities to [0, 1]:
normalized_brains = brains/1328
And then by using the binomial distribution to binarize each pixel:
binarized_brains = np.random.binomial(1, (normalized_brains))
The plotted result looks correct:

A 0 pixel intensity represents black (background) and 1 pixel intensity represents white (brain).
I experimented by implementing another method to normalise an image from this post but it gave me just a black image. This is because np.finfo(np.float64) is 1.7976931348623157e+308, so the normalization step
normalized_brains = brains/1.7976931348623157e+308
just returned an array of zeros which in the binarizition step also led to an array of zeros.
Am I binarising my images using a correct method?
Your method of converting the image to a binary image basically amounts to random dithering, which is a poor method of creating the illusion of grey values on a binary medium. Old-fashioned print is a binary medium, they have fine-tuned the methods to represent grey-value photographs in print over centuries. This process is called halftoning, and is shaped in part by properties of ink on paper, that we do not have to deal with in binary images.
So what methods have people come up with outside of print? Ordered dithering (mostly Bayer matrix), and error diffusion dithering. Read more about dithering on Wikipedia. I wrote a blog post showing how to implement all of these methods in MATLAB some years ago.
I would recommend you use error diffusion dithering for your particular application. Here is some code in MATLAB (taken from my blog post liked above) for the Floyd-Steinberg algorithm, I hope that you can translate this to Python:
img = imread('https://i.stack.imgur.com/d5E9i.png');
img = img(:,:,1);
out = double(img);
sz = size(out);
for ii=1:sz(1)
for jj=1:sz(2)
old = out(ii,jj);
%new = 255*(old >= 128); % Original Floyd-Steinberg
new = 255*(old >= 128+(rand-0.5)*100); % Simple improvement
out(ii,jj) = new;
err = new-old;
if jj<sz(2)
% right
out(ii ,jj+1) = out(ii ,jj+1)-err*(7/16);
end
if ii<sz(1)
if jj<sz(2)
% right-down
out(ii+1,jj+1) = out(ii+1,jj+1)-err*(1/16);
end
% down
out(ii+1,jj ) = out(ii+1,jj )-err*(5/16);
if jj>1
% left-down
out(ii+1,jj-1) = out(ii+1,jj-1)-err*(3/16);
end
end
end
end
imshow(out)

Resampling the image before applying the dithering greatly improves the results:
img = imresize(img,4);
% (repeat code above)
imshow(out)

NOTE that the above process expects the input to be in the range [0,255]. It is easy to adapt to a different range, say [0,1328] or [0,1], but it is also easy to scale your images to the [0,255] range.
Have you tried a threshold on the image?
This is a common way to binarize images, rather than trying to apply a random binomial distribution. You could try something like:
binarized_brains = (brains > threshold_value).astype(int)
which returns an array of 0s and 1s according to whether the image value was less than or greater than your chosen threshold value.
You will have to experiment with the threshold value to find the best one for your images, but it does not need to be normalized first.
If this doesn't work well, you can also experiment with the thresholding options available in the skimage filters package.
IT is easy in OpenCV. as mentioned a very common way is defining a threshold, But your result looks like you are allocating random values to your intensities instead of thresholding it.
import cv2
im = cv2.imread('brain.png', cv2.CV_LOAD_IMAGE_GRAYSCALE)
(th, brain_bw) = cv2.threshold(imy, 128, 255, cv2.THRESH_BINARY | cv2.THRESH_OTSU)
th = (DEFINE HERE)
im_bin = cv2.threshold(im, th, 255, cv
cv2.imwrite('binBrain.png', brain_bw)
brain
binBrain